Chris Eliasmith: We Have Not Yet Learned What The Brain Has To Teach Us!
Socrates / Podcasts
Posted on: January 21, 2013 / Last Modified: October 14, 2021
Podcast: Play in new window | Download | Embed
Subscribe: RSS
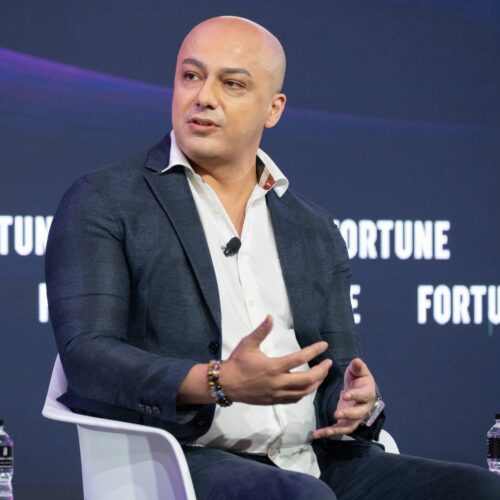
Prof. Chris Eliasmith is currently the director of the Centre for Theoretical Neuroscience at the University of Waterloo and the team leader behind SPAUN – the brain simulation project that recently made news around the world. So, when I discovered that Eliasmith’s lab is just over an hour worth of driving from my place, I decided that I would take this opportunity to go talk to him in person.
During my Singularity 1 on 1 interview with Chris, we discuss a variety of topics such as the story behind his desire to create a whole-brain simulation; SPAUN (the Semantic Pointer Architecture Unified Network), and the hardware requirements to run it; whether SPAUN has thoughts and feelings and how would we know if it did; the ethical issues behind creating a brain-in-a-vat AI; the relationship between philosophy and engineering; his upcoming book How To Build A Brain; Eliasmith’s thoughts on Deep Blue, Watson, Blue Brain, SyNAPSE and Ray Kurzweil‘s How To Create A Mind; his take on the technological singularity…
My favorite quote from Prof. Chris Eliasmith is:
We have not yet learned what the brain has to teach us!
As always you can listen to or download the audio file above or scroll down and watch the video interview in full. To show your support you can write a review on iTunes, make a direct donation, or become a patron on Patreon.
What is SPAUN?
You can download and test the brain simulation yourself at nengo.ca
Who is Chris Eliasmith?
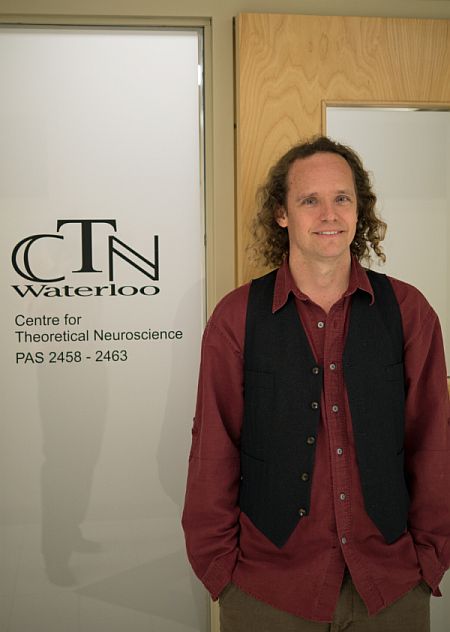 Dr. Eliasmith is currently the director of the Centre for Theoretical Neuroscience at Waterloo. The Centre focuses on mathematical characterizations of a variety neural systems, from individual ion channels to large-scale networks. Dr. Eliasmith is also head of the Computational Neuroscience Research Group (CNRG) at the Centre. This group is developing and applying a general framework for modeling the function of complex neural systems (the Neural Engineering Framework or NEF). The NEF is grounded in the principles of signal processing, control theory, statistical inference, and good engineering design. It provides a rational and robust strategy for simulating and evaluating the function of a wide variety of specific biological neural circuits. Members of the group applied the NEF to projects characterizing sensory processing, motor control, and cognitive function.
Dr. Eliasmith is currently the director of the Centre for Theoretical Neuroscience at Waterloo. The Centre focuses on mathematical characterizations of a variety neural systems, from individual ion channels to large-scale networks. Dr. Eliasmith is also head of the Computational Neuroscience Research Group (CNRG) at the Centre. This group is developing and applying a general framework for modeling the function of complex neural systems (the Neural Engineering Framework or NEF). The NEF is grounded in the principles of signal processing, control theory, statistical inference, and good engineering design. It provides a rational and robust strategy for simulating and evaluating the function of a wide variety of specific biological neural circuits. Members of the group applied the NEF to projects characterizing sensory processing, motor control, and cognitive function.
Work at the CNRG divides into two main areas: applications and theoretical development. Theoretical work includes extending the NEF to be more general (e.g. account for a wider range of single cell dynamics), more biologically plausible (e.g., capture network physiology and topology more precisely), and more adaptive (e.g., including better adaptive filtering, learning, etc.). In addition, the CNRG members are exploring general principles for brain function to explain not only how neural systems implement complex dynamics (the focus of the NEF), but also what neural systems are designed to do in general – i.e., what the basic functional principles of the brain are.
Applications consist of building complex networks of single cells to test hypotheses about the functioning of a given neural system. The results of such simulations are compared against available neural data, and used to make novel predictions. CNRG members have constructed models of working memory, locomotion, decision making, posture control, the basal ganglia (implicated in Parkinsons Disease), rodent navigation, and language use, among others.
Some members of the CNRG have recently begun developing applications of related principles to problems in machine intelligence. Specifically, they are constructing novel methods for automatic text understanding that can be used to support classification and clustering. The focus of this work is on integrating semantics and structure in an appropriate flat vector representation.